Annotation Process
Two trained annotators, a data scientist (A1) and a post-doc researcher (A2), conducted the annotation process on nuclei segmentation by an expert pathologist to generate the ground truth masks. The annotation protocol was inspired by the CryoNuSeg and MoNuSeg datasets protocols, which was further enhanced by adding a first validation using the masks generated automatically by the FIJI ImageJ software. The annotation process:
- Generate unsupervised automatic nuclei semantic segmentation masks using
the Fiji ImageJ software, which serve as preliminary annotations for the
tissue slides. These annotations provide a rough estimate of the nuclei locations, simplifying the initial annotation process for the two annotators A1
& A2.
- Perform manual nuclei semantic segmentation using the LabelStudio segmentation tool by the two annotators A1 & A2.
- Conduct a second round of intra-observation by A2 to validate the annotations made by A1.
- Validate the final manual-generated ground truth masks by three expert
pathologists with 4, 10 and 15 years of experience.
- Correct the manual nuclei annotations based on the markups and observations of the three expert pathologists.
- Conduct a final validation by three expert pathologists to ensure that the
annotations meet the highest standards to help the pathologist make the
diagnosis
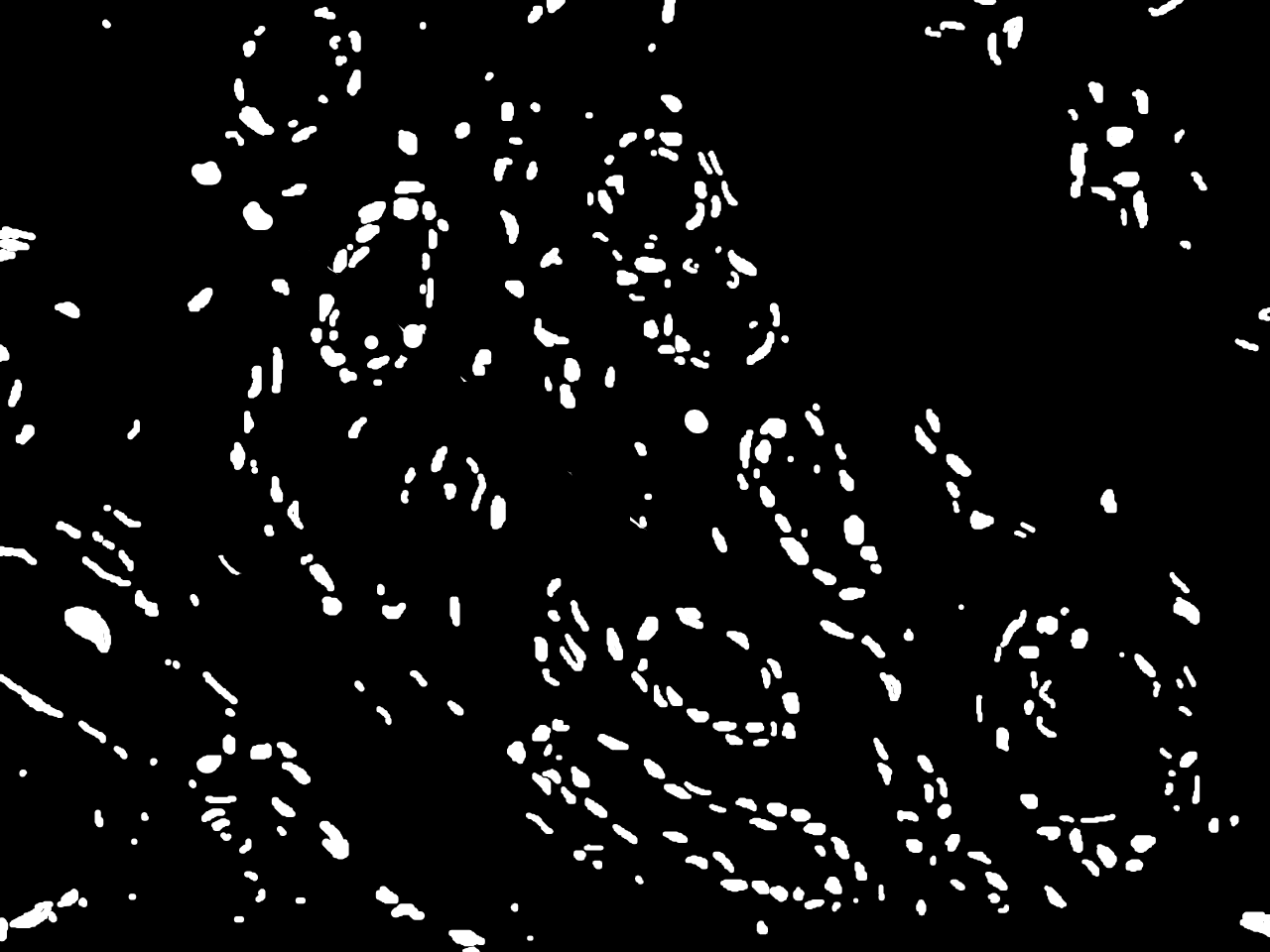
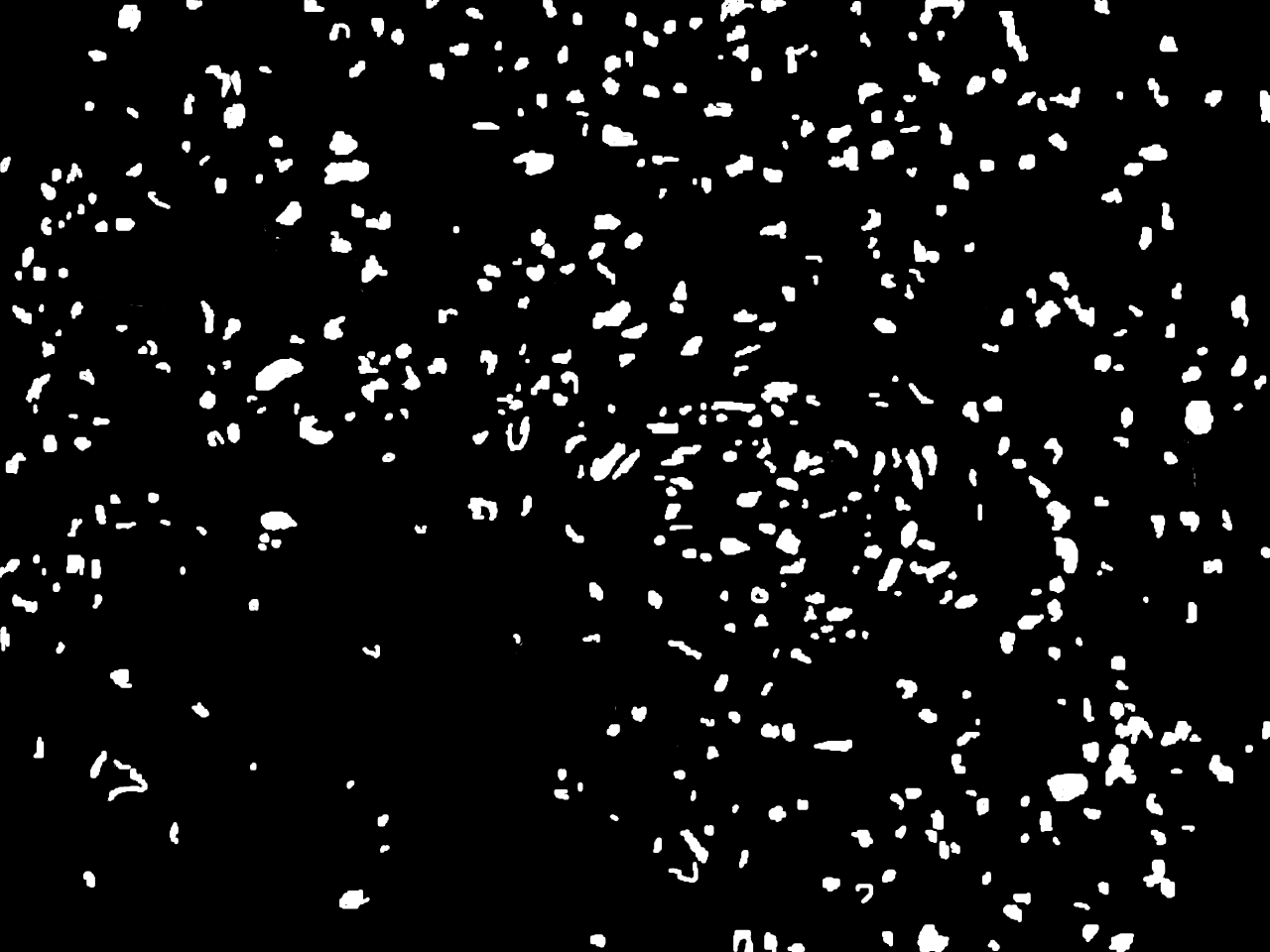
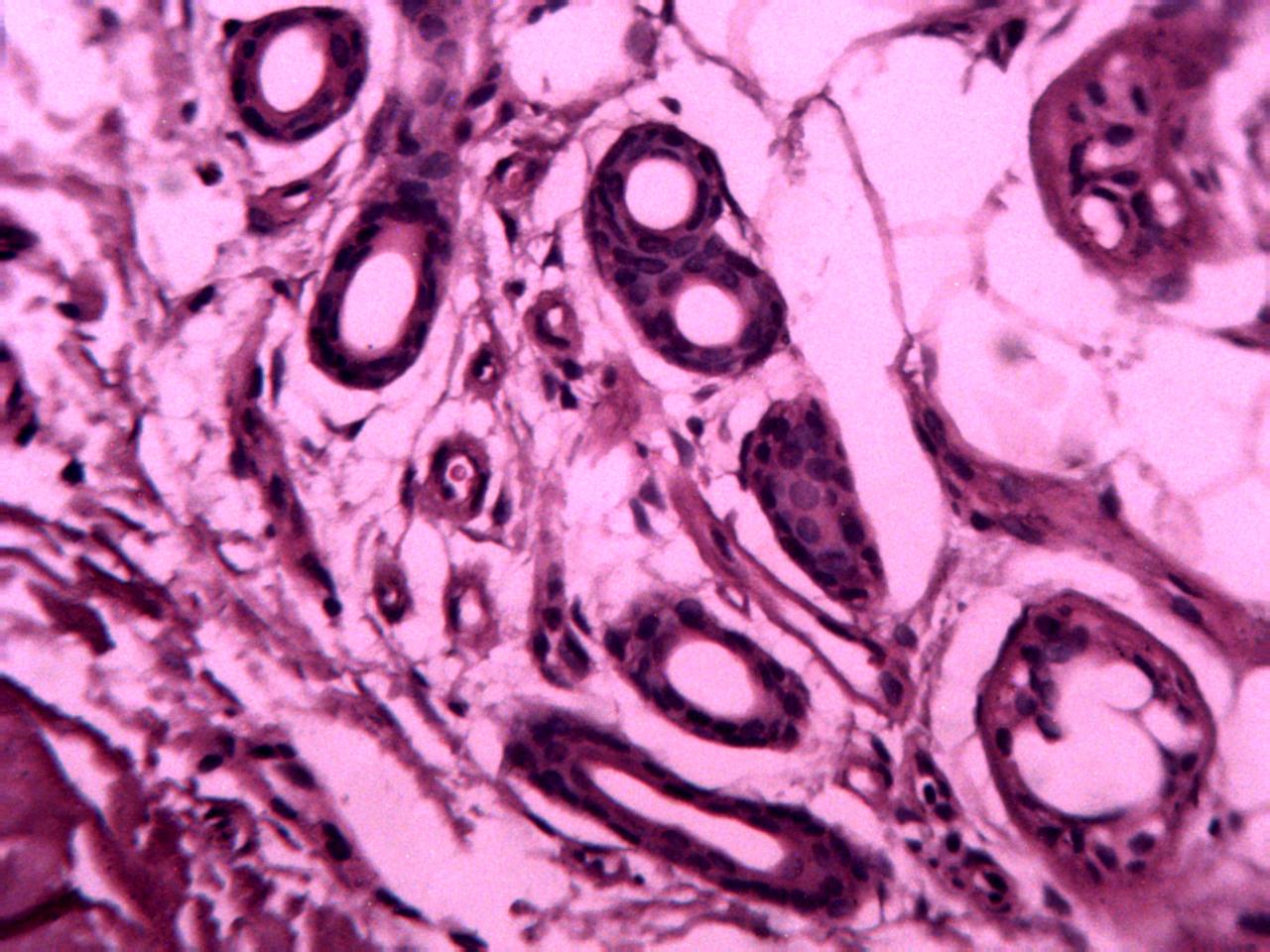
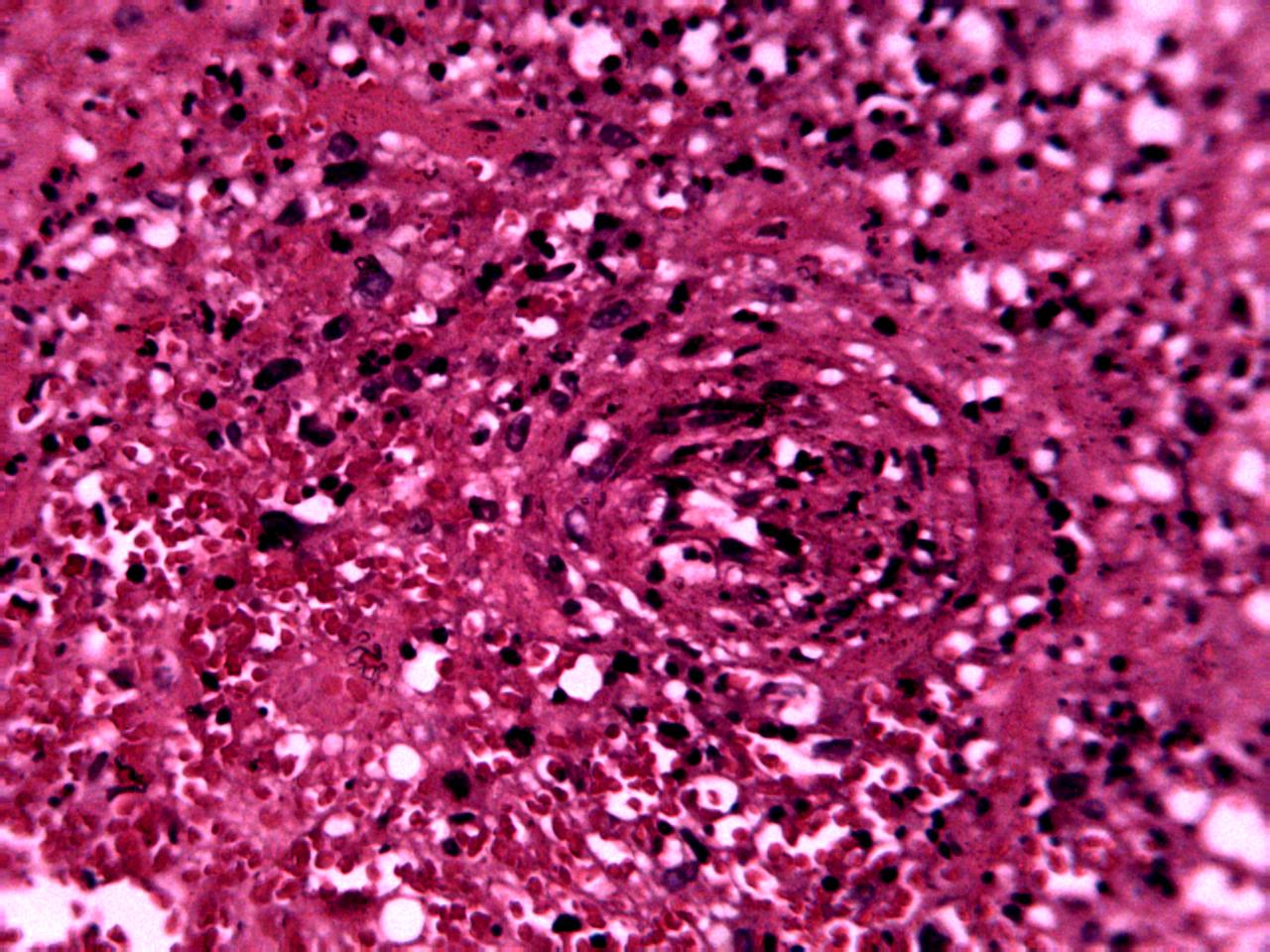
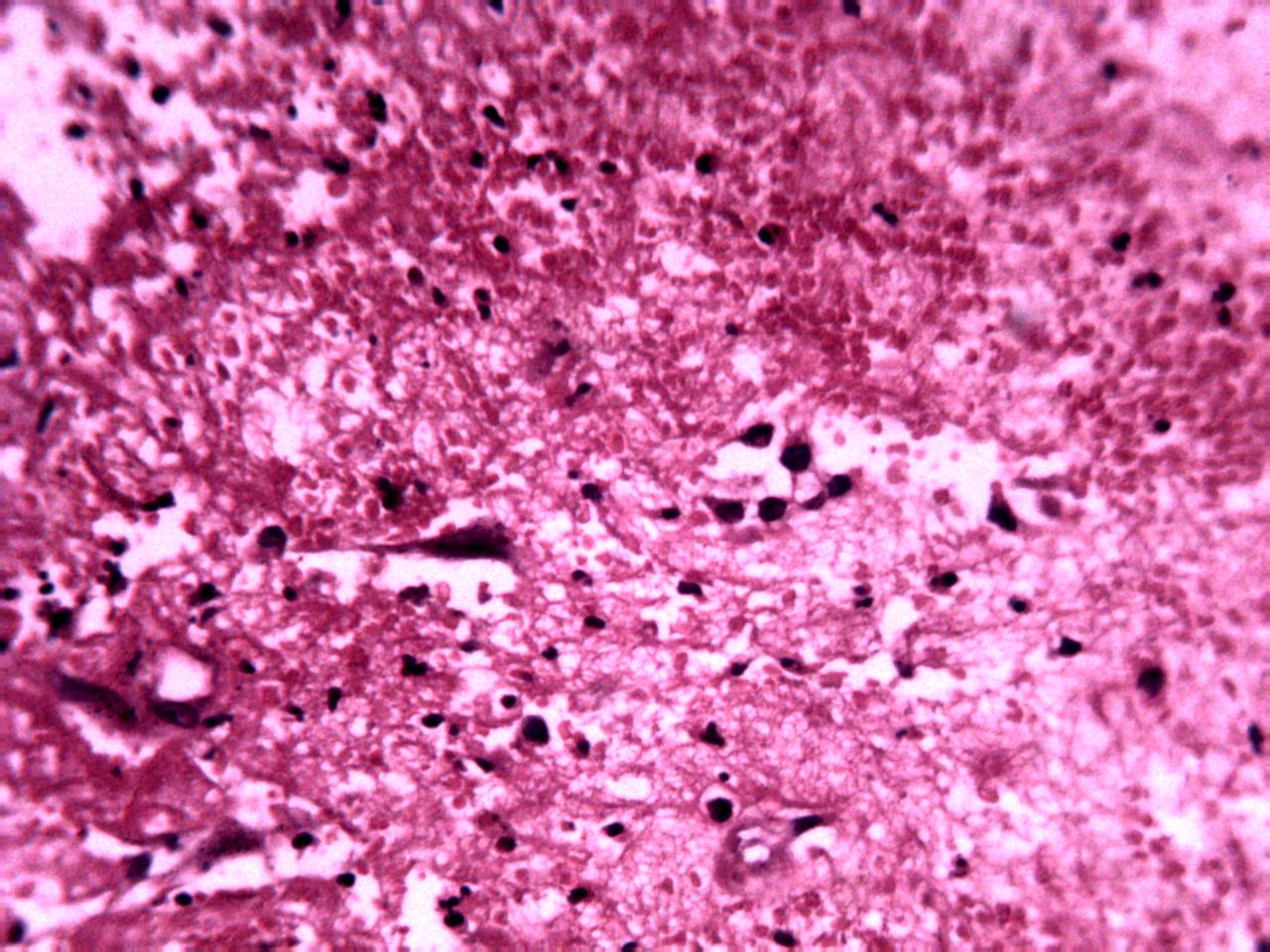
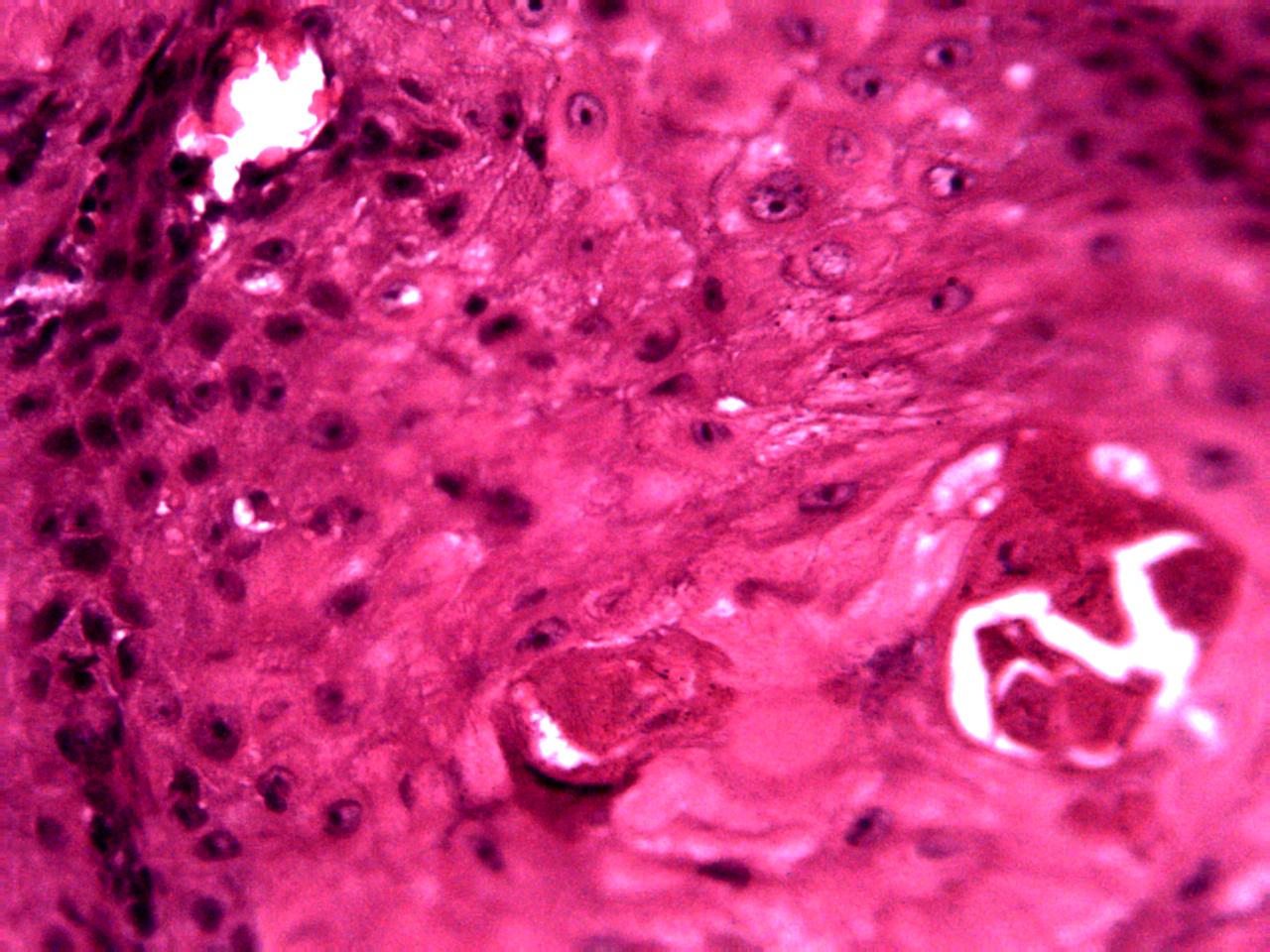
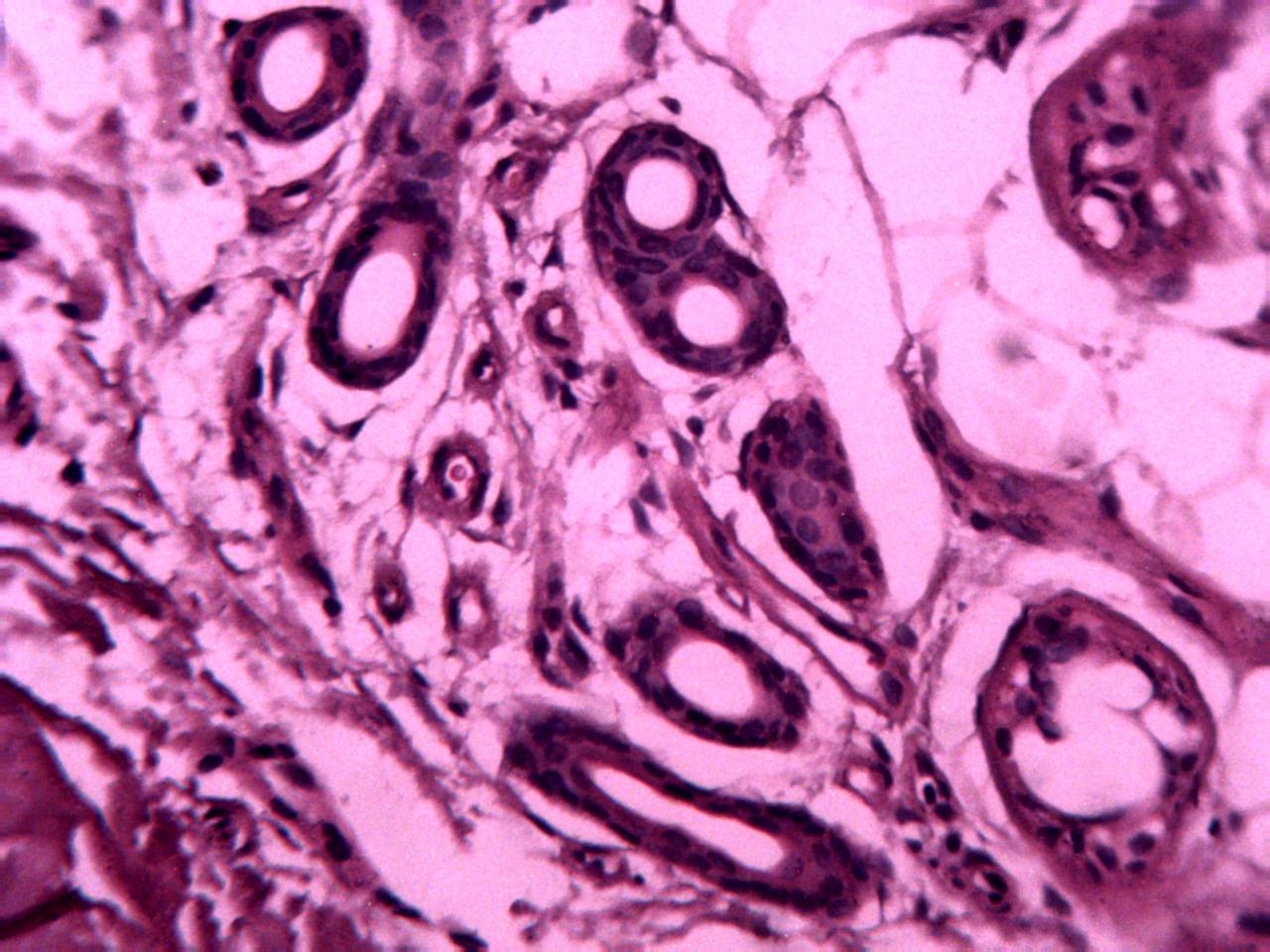
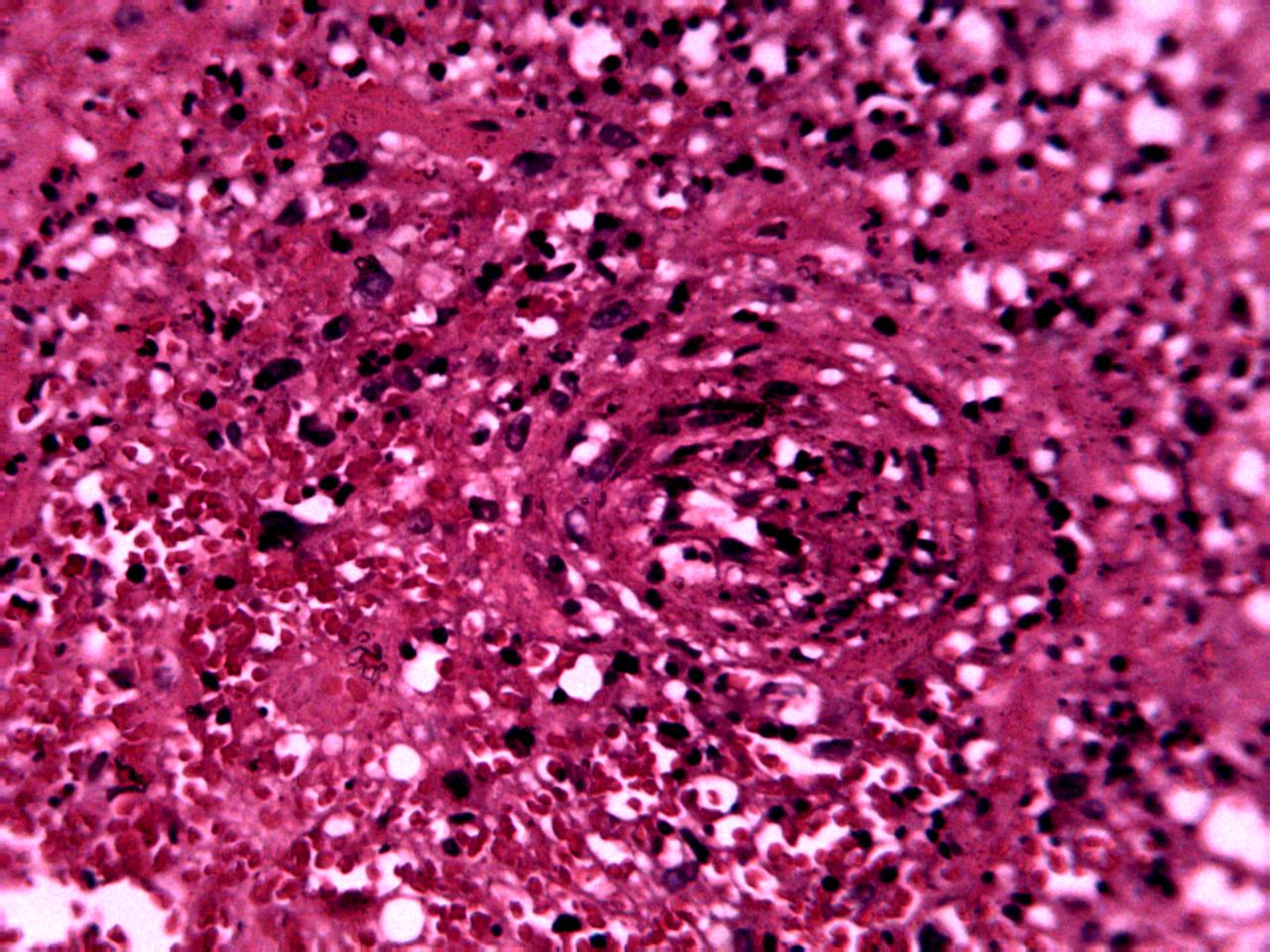
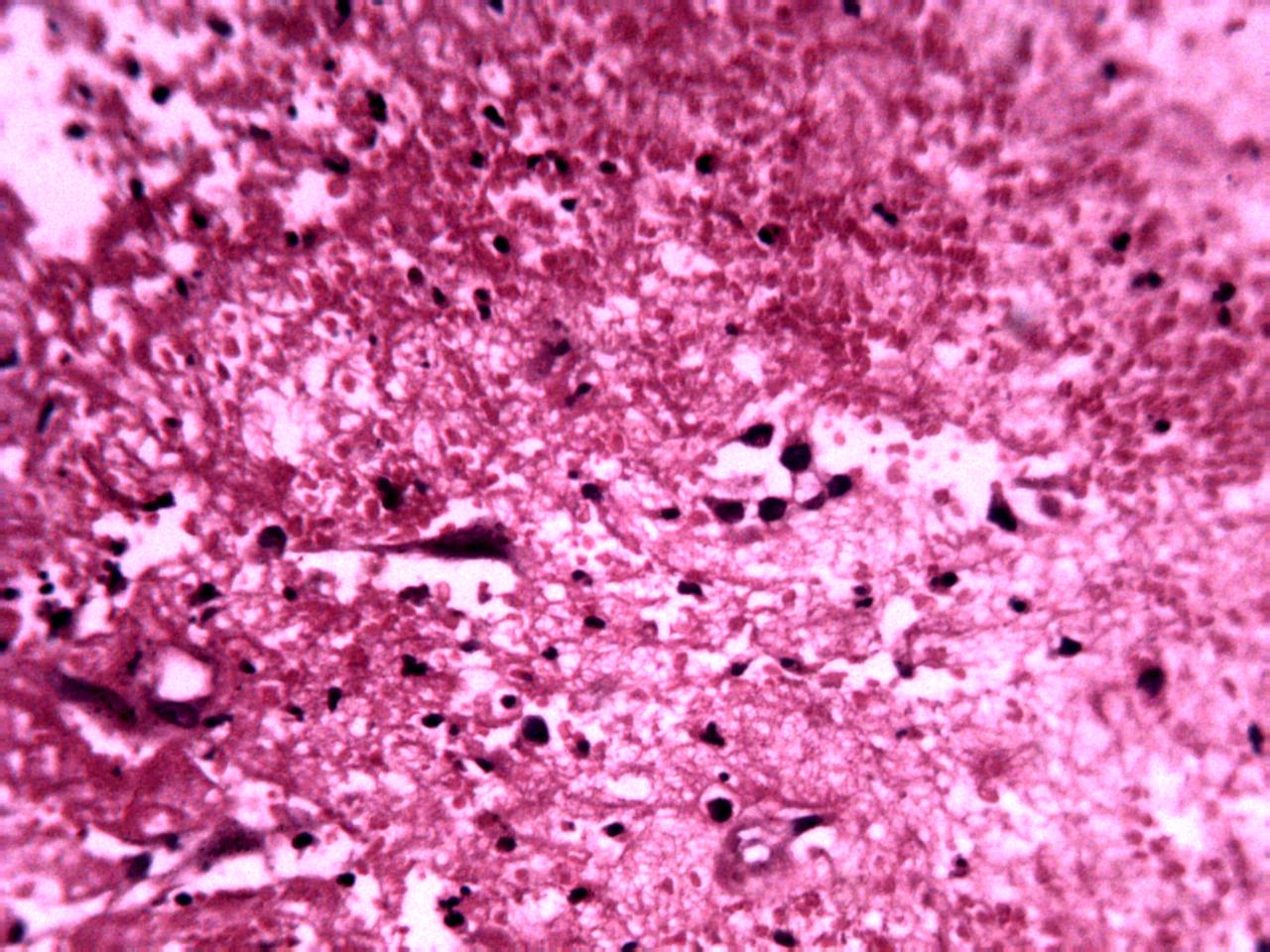
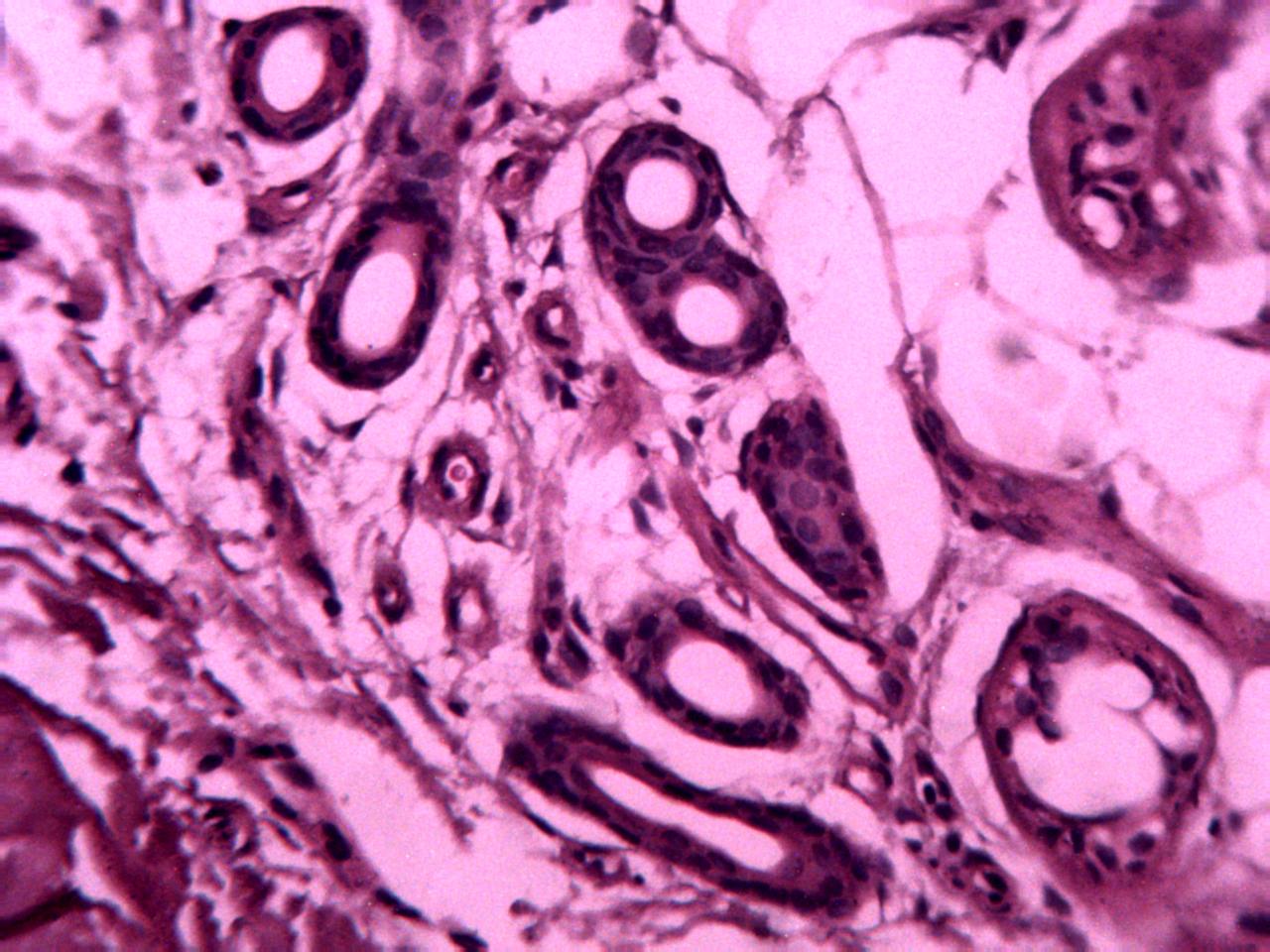
Annotations curated by a certified pathologist from Nigeria (Ahmado Bello University)
Samples of breast, skin, cervical and ingunal and their annotation masks validated by a certified pathologist from Nigeria
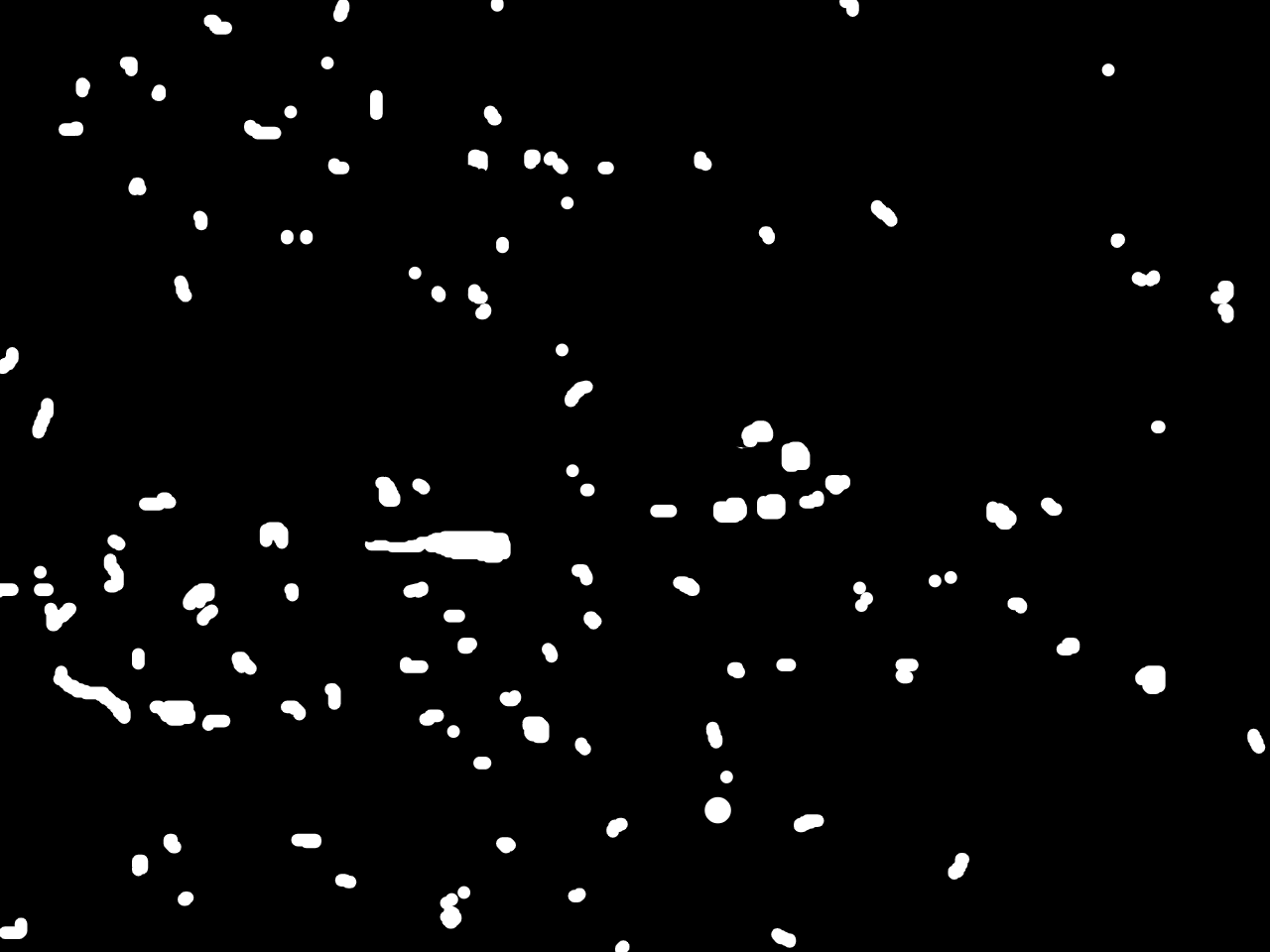
Annotations curated by a certified pathologist from Morocco (Amana Laboratory)
Samples of breast, skin, cervical and ingunal and their annotation masks validated by a certified pathologist from Morocco
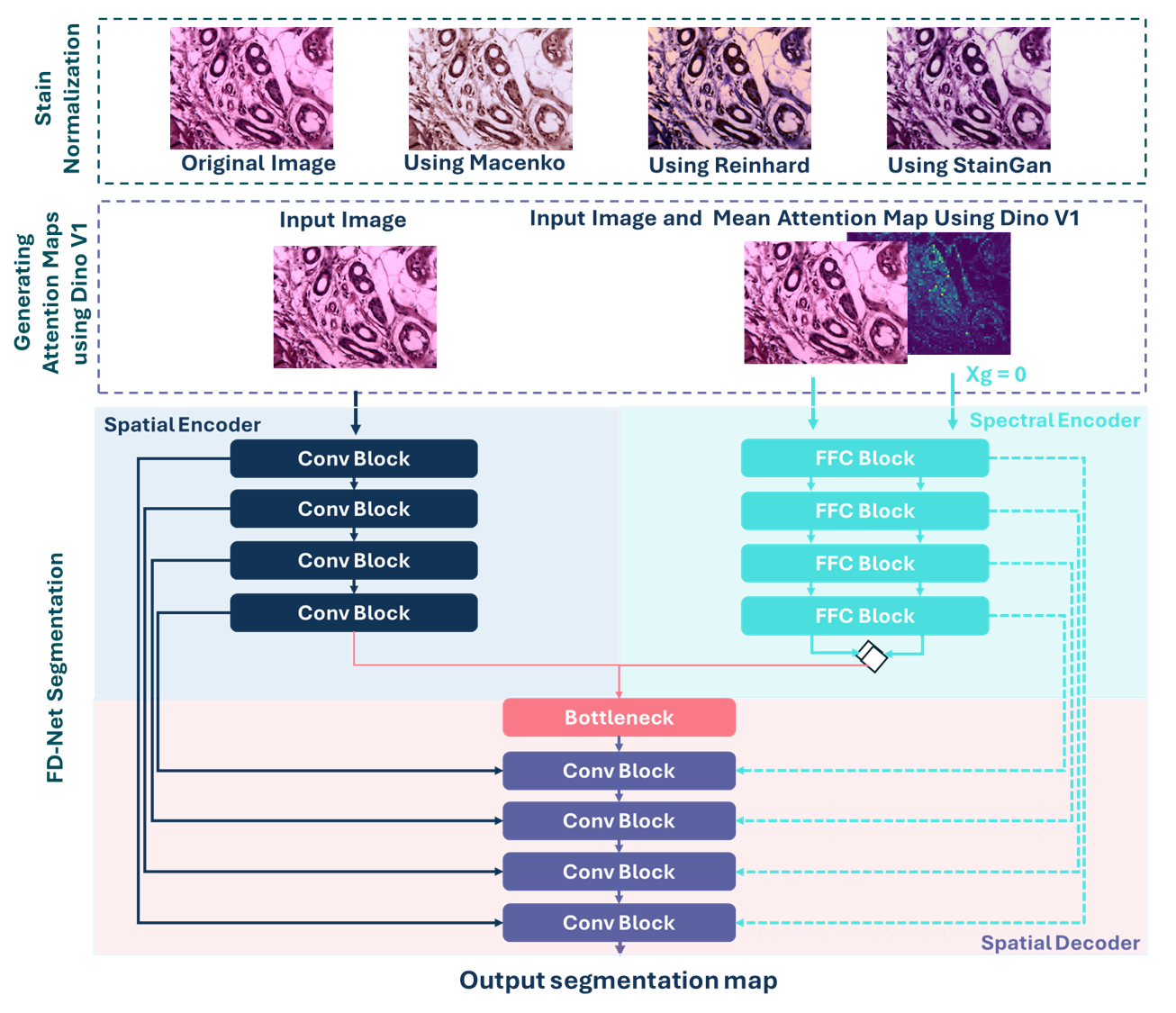
FD-NET
The FD-Net model consists of Two Branches:
- Spatial encoder branch that processes spatial features and consists of convolutional blocks.
- Spectral branch that processes spectral features and consists of fast Fourier convolutional blocks.
- For the spatial encoder branch: The input image can be either the original image or a stain-normalized preprocessed image using Macenko, Reinhard, or StainGAN methods.
- For the spectral encoder branch: The input image and the generated mean attention map using Dino v1 are fed to the spectral branch.
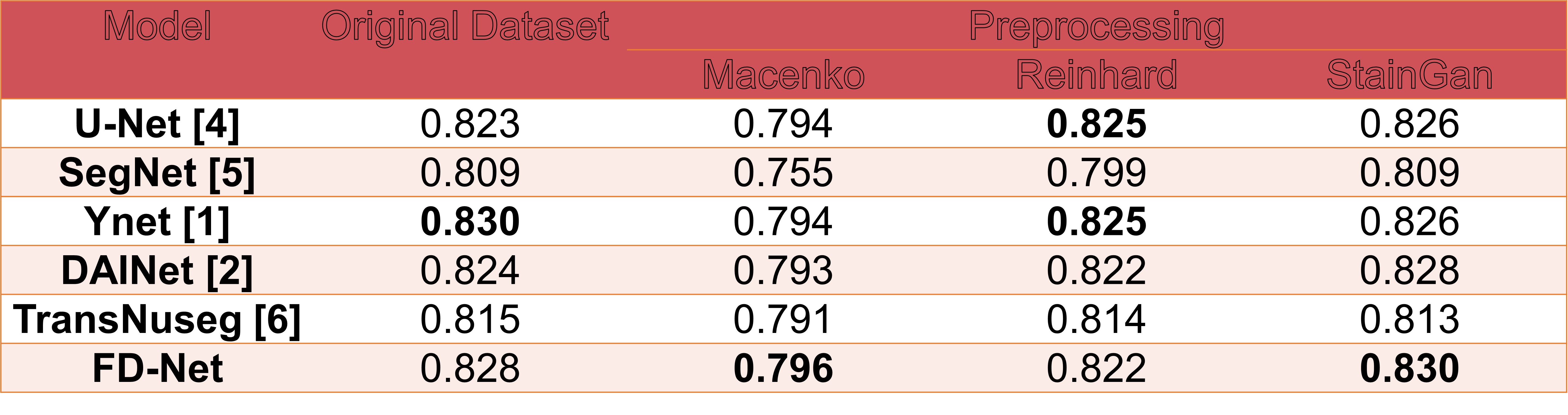
Results in terms of average Dice Score
The best performance achieved a higher average Dice score of 0.830 using both Y-Net with the original AMONuSeg and FD-Net with the StainGAN pre-processed dataset.
BibTeX
@InProceedings{Zer_AMONuSeg_MICCAI2024,
author = { Zerouaoui, Hasnae and Oderinde, Gbenga Peter and Lefdali, Rida and Echihabi, Karima and Akpulu, Stephen Peter and Agbon, Nosereme Abel and Musa, Abraham Sunday and Yeganeh, Yousef and Farshad, Azade and Navab, Nassir},
title = { { AMONuSeg: A Histological Dataset for African Multi-Organ Nuclei Semantic Segmentation } },
booktitle = {proceedings of Medical Image Computing and Computer Assisted Intervention -- MICCAI 2024},
year = {2024},
publisher = {Springer Nature Switzerland},
volume = {LNCS 15009},
month = {October},
page = {pending}
}